Document Portal
One Solution for all your Documents
Pydio Cells is a document sharing and collaboration platform that puts you in full control of your documents in a way Saas solutions can’t - combining fast performance, huge file transfer sizes, granular security, and advanced workflow automations in a single platform.
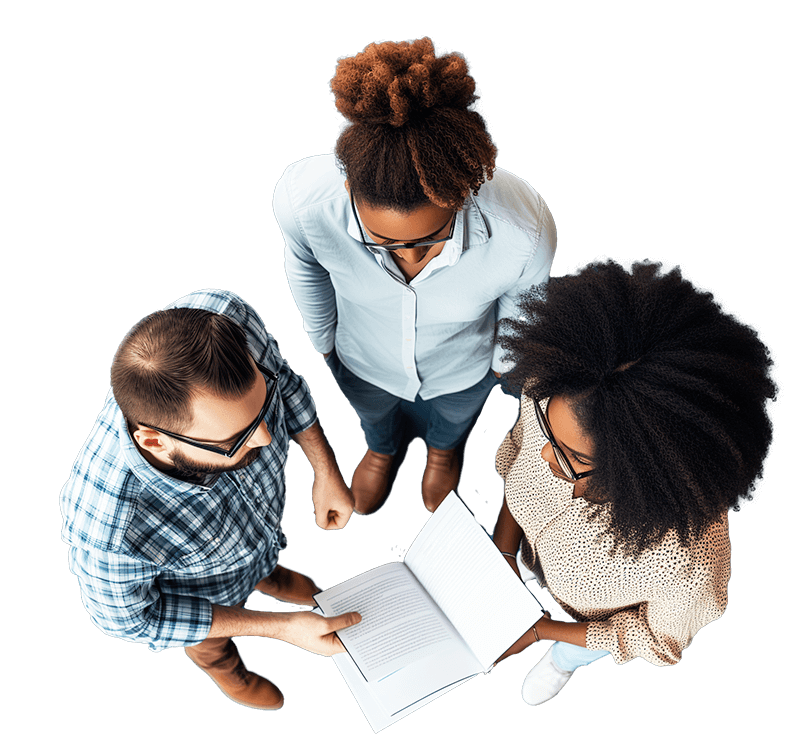

6 Reasons Pydio May Be the Platform You Need
We Need Complete Control of our Docs
We Need to Stop Platform Creep
We need a Cloud-Ready Solution
We Need to Keep our Documents Secure
We Need a Platform that’s Easy and Intuitive
We Need to Automate Right Now!
Collaboration Powered By Pydio



Documents Made Simple
Designed For Teams
Your sharing and collaboration platform should enable your teams to harness their ideas and creativity, not create roadblocks and hold them back.
Secured For Organizations
SaaS solutions put your data security in someone else’s hands. Cells is self-hosted sharing & collaboration without trade-offs.
Optimized For IT
Forget what you’ve heard about hard-to-manage open-core software. Pydio is designed to meet the needs of even the most demanding IT departments.
Automate. Accelerate. Connect.
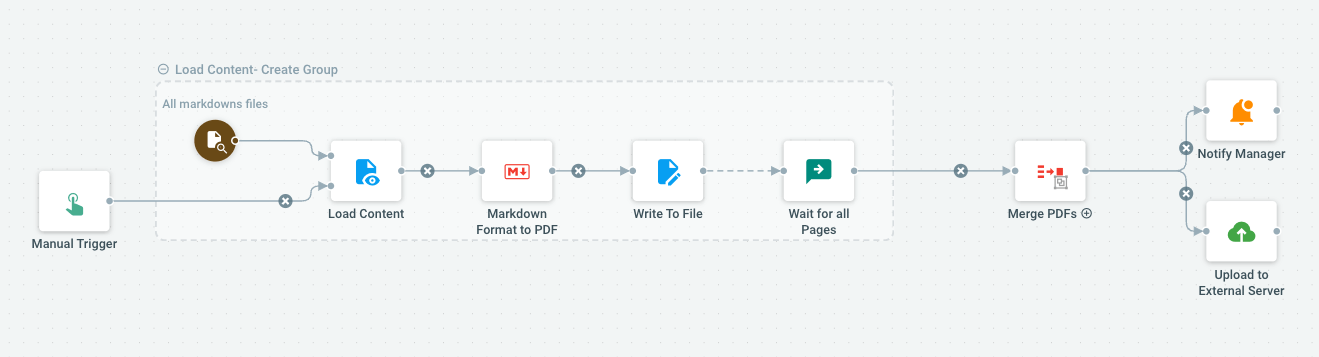
A Single Platform for All Your Document Needs
File Sharing & Collaboration
Large File Transfers
Virtual Data Room
Digital Asset Management
Document Management
Knowledge Management
Document Portal
Remote Is the New Normal
What Customers Say About Pydio
Dimitri R - Security & System administrator at major Insurance company
Jeremy Godefroy - Deputy IT Manager at Inter Public Group
Baptiste Mary - Computer Science and Digital at CY Cergy Paris Université